Exporting rtables Tables to PDF
v06-rtables.RmdThis vignette covers export_tfl() as used with rtables
VTableTree objects. For data-frame tables built with
tfl_table(), see
vignette("v02-tfl_table_intro"). For gt tables, see
vignette("v05-gt_tables"). For figure output, see
vignette("v01-figure_output").
library(writetfl)
library(rtables)
#> Loading required package: formatters
#>
#> Attaching package: 'formatters'
#> The following object is masked from 'package:base':
#>
#> %||%
#> Loading required package: magrittr
#>
#> Attaching package: 'rtables'
#> The following object is masked from 'package:utils':
#>
#> str
library(grid)Basic usage
Pass a VTableTree object directly to
export_tfl(). The main title, subtitles, main footer, and
provenance footer are automatically extracted and placed in writetfl’s
annotation zones (caption and footnote), while the table body is
rendered as monospace text via toString() and wrapped in a
grid textGrob.
lyt <- basic_table(
title = "Iris Sepal Length by Species",
subtitles = "Mean values",
main_footer = "Source: Anderson (1935)."
) |>
split_cols_by("Species") |>
analyze("Sepal.Length", mean)
tbl <- build_table(lyt, iris)
export_tfl(tbl, preview = TRUE)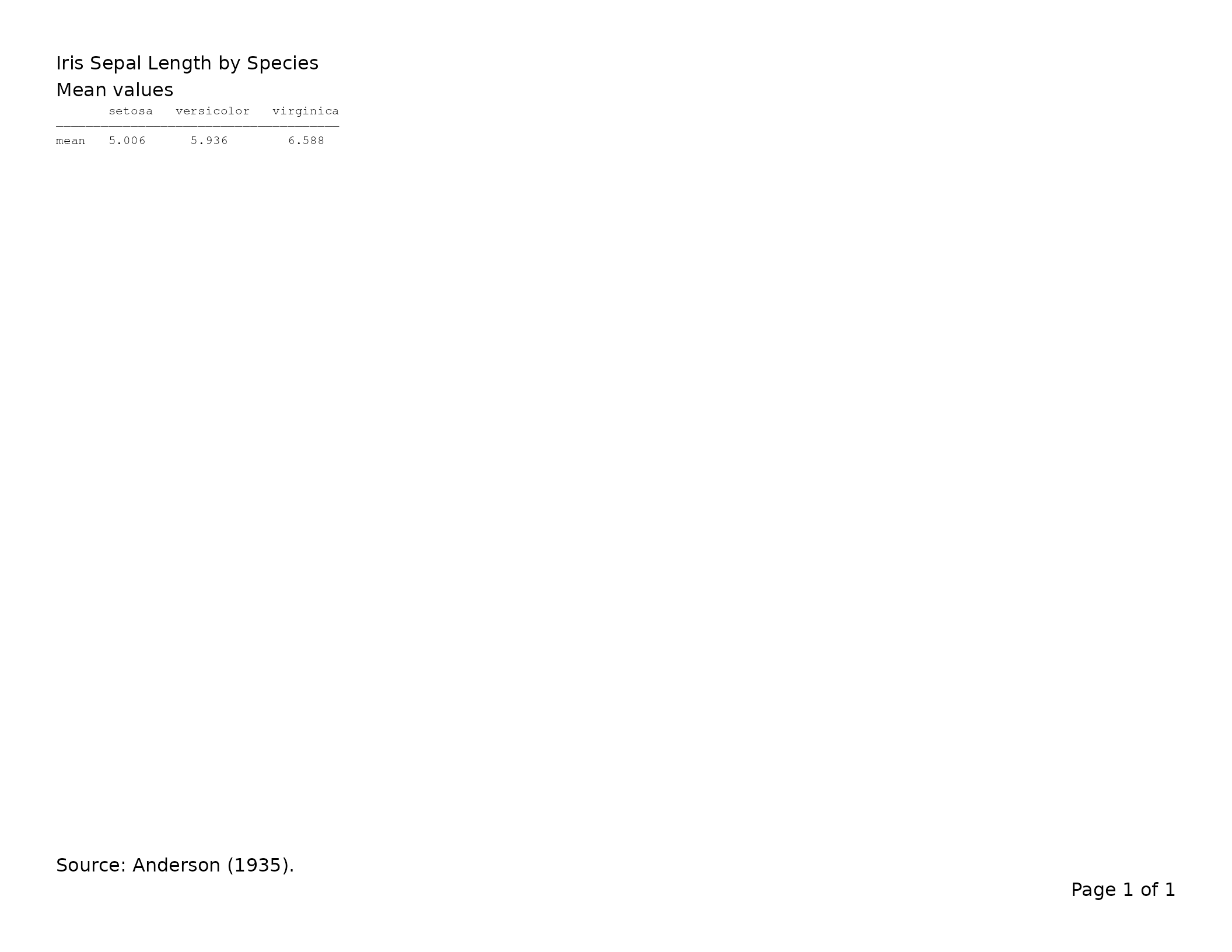
Why annotations are extracted
rtables normally renders its title, subtitles, and footers as part of
the text output. When placed inside writetfl’s page layout, this would
cause duplication — the annotations would appear both in the text
rendering and in writetfl’s header/footer zones. To avoid this,
export_tfl() extracts rtables annotations into writetfl’s
annotation fields and strips them from the rtables object before
rendering.
The mapping is:
| rtables annotation | writetfl field |
|---|---|
main_title |
caption (first line) |
subtitles |
caption (subsequent lines, joined with
\n) |
main_footer |
footnote |
prov_footer |
footnote (appended after main footer) |
Adding page layout elements
All of writetfl’s page layout arguments work with rtables tables.
Pass them via ... just as you would for figures.
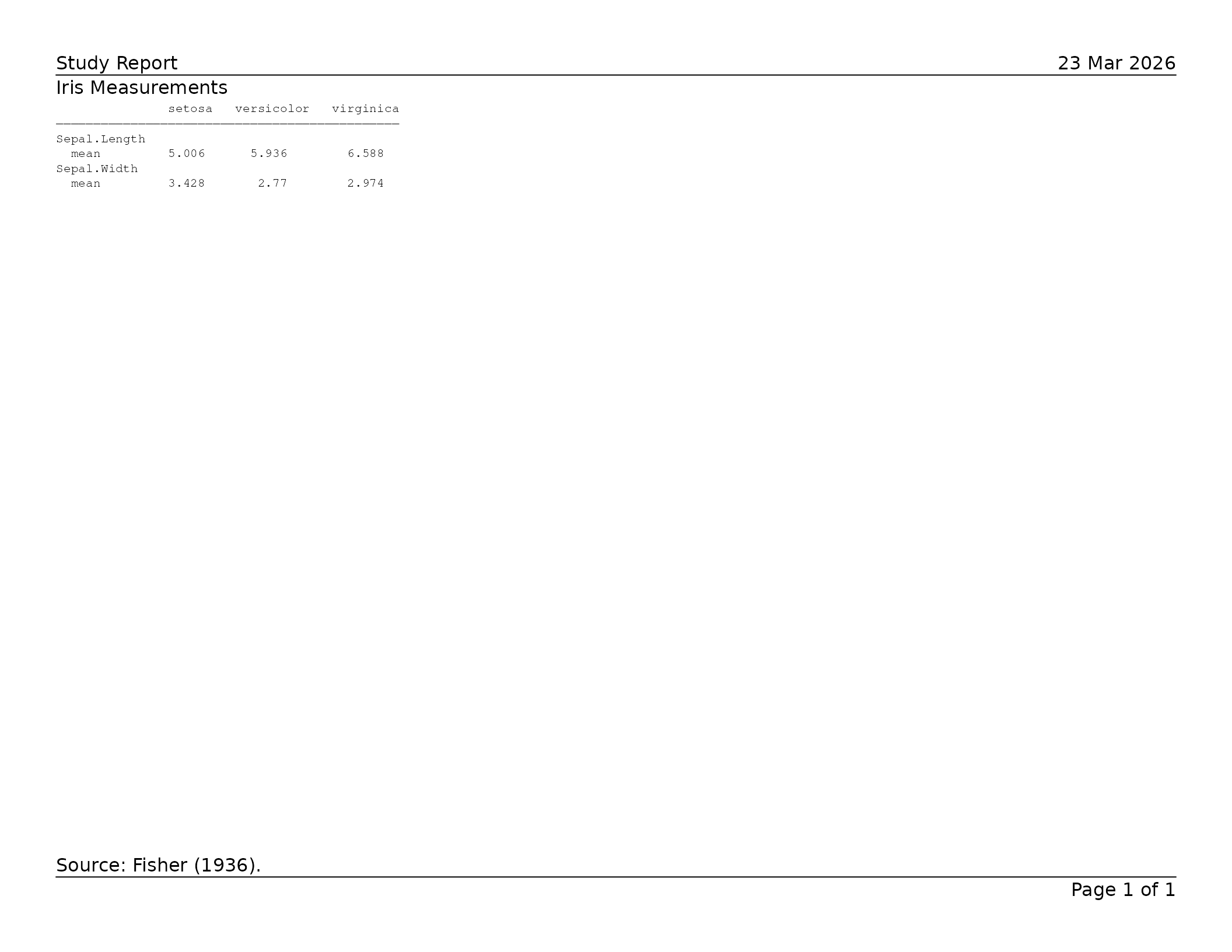
lyt <- basic_table(
title = "Iris Measurements",
main_footer = "Source: Fisher (1936)."
) |>
split_cols_by("Species") |>
analyze(c("Sepal.Length", "Sepal.Width"), mean)
tbl <- build_table(lyt, iris)
export_tfl(
tbl,
preview = TRUE,
header_left = "Study Report",
header_right = format(Sys.Date(), "%d %b %Y"),
header_rule = TRUE,
footer_rule = TRUE
)
Multiple rtables tables
Pass a list of VTableTree objects to produce a
multi-page PDF with one table per page. Each table’s annotations are
extracted independently.
tbl1 <- basic_table(title = "Table 1: Sepal Length") |>
split_cols_by("Species") |>
analyze("Sepal.Length", mean) |>
build_table(iris)
tbl2 <- basic_table(title = "Table 2: Petal Length") |>
split_cols_by("Species") |>
analyze("Petal.Length", mean) |>
build_table(iris)
export_tfl(
list(tbl1, tbl2),
file = "two-tables.pdf",
header_left = "Appendix",
header_rule = TRUE
)Automatic pagination
When an rtables table is too tall to fit on a single page,
export_tfl() uses rtables’ built-in
paginate_table() to split it across pages. Row group
boundaries are respected — a group is never split across pages.
big_data <- data.frame(
arm = rep(c("Treatment", "Control"), each = 50),
site = rep(paste0("Site ", 1:10), each = 10),
outcome = rnorm(100, 50, 10)
)
lyt <- basic_table(
title = "Subject Outcomes by Site",
main_footer = "Source: Clinical Trial XY-001."
) |>
split_cols_by("arm") |>
split_rows_by("site") |>
analyze("outcome", mean)
tbl <- build_table(lyt, big_data)
export_tfl(
tbl,
file = "paginated.pdf",
header_left = "Study Report",
header_rule = TRUE,
footer_rule = TRUE
)Each page carries the same caption and footnote from the original rtables object.
How pagination works
- The available content height is measured (page height minus margins, headers, footers, caption, and footnote).
- Content dimensions are converted to lines-per-page
(
lpp) and characters-per-page (cpp) using font metrics. -
rtables::paginate_table()splits the table, respecting row group boundaries and rtables’ own split rules. - Each page is rendered to text via
toString()and wrapped in atextGrobwith monospace font.
Font control
The rendering font can be controlled via ...
parameters:
export_tfl(
tbl,
file = "custom-font.pdf",
rtables_font_family = "Courier",
rtables_font_size = 10,
rtables_lineheight = 1.2
)Preserved rtables features
The following rtables features are preserved through the
toString() rendering pipeline:
| Feature | Preserved? | Notes |
|---|---|---|
main_title |
Yes | Extracted as writetfl caption |
subtitles |
Yes | Extracted as writetfl caption |
main_footer |
Yes | Extracted as writetfl footnote |
prov_footer |
Yes | Extracted as writetfl footnote |
split_cols_by() |
Yes | Column structure rendered by toString |
split_rows_by() |
Yes | Row groups with nesting and indentation |
analyze() |
Yes | Analysis rows with formatting |
summarize_row_groups() |
Yes | Group summary rows |
add_colcounts() |
Yes | Column N counts |
append_topleft() |
Yes | Top-left corner label |
tab_fn_*() footnotes |
Yes | Referential footnotes |
| Section dividers | Yes |
horizontal_sep, section_div
|
| Cell formatting | Yes | All rcell() format strings |
Column splits
lyt <- basic_table(title = "Column Split Example") |>
split_cols_by("Species") |>
analyze(c("Sepal.Length", "Sepal.Width", "Petal.Length", "Petal.Width"),
mean)
tbl <- build_table(lyt, iris)
export_tfl(tbl, preview = TRUE)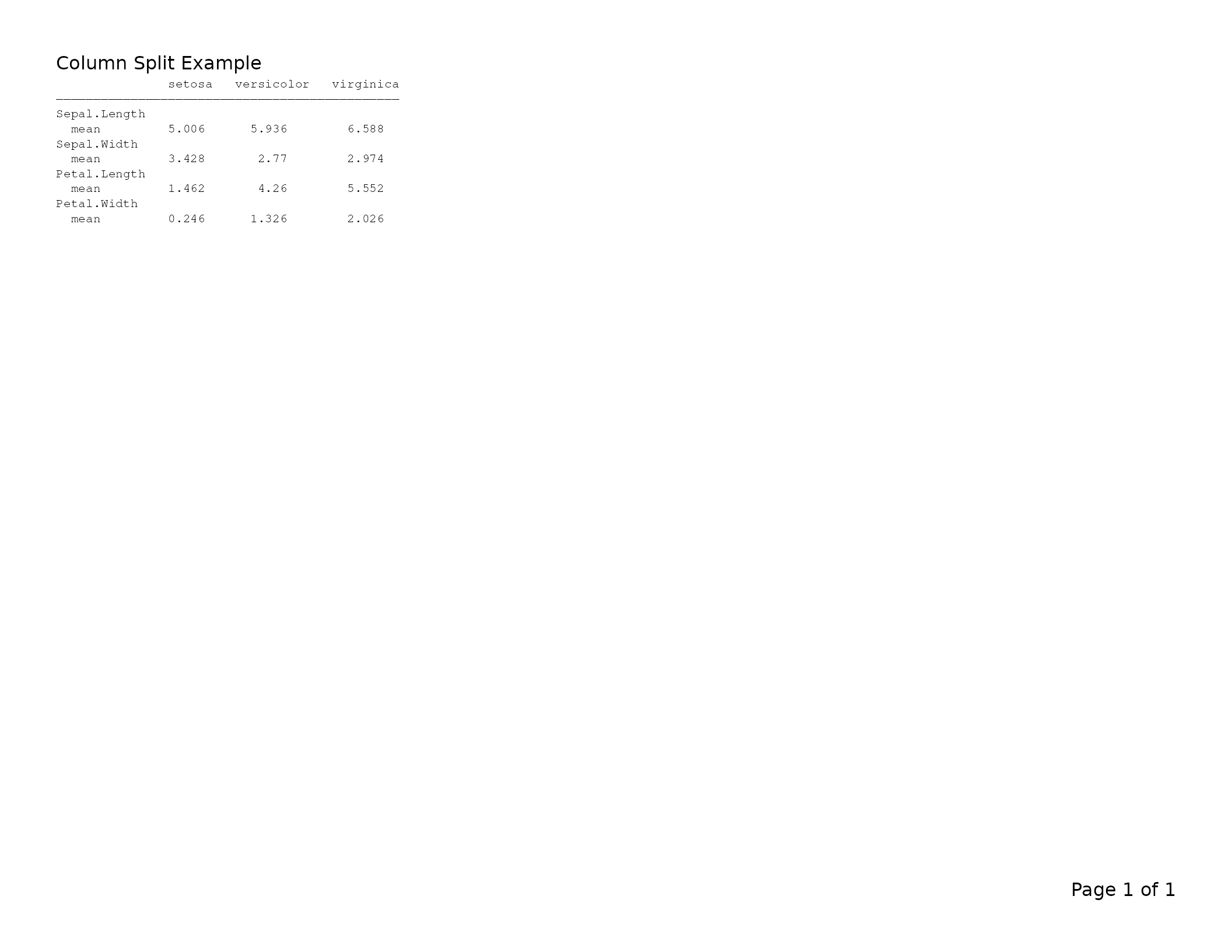
Row groups with nesting
lyt <- basic_table(title = "Nested Row Groups") |>
split_rows_by("Species") |>
analyze(c("Sepal.Length", "Petal.Length"), mean)
tbl <- build_table(lyt, iris)
export_tfl(tbl, preview = TRUE)
Column counts
lyt <- basic_table(title = "With Column Counts") |>
split_cols_by("Species") |>
add_colcounts() |>
analyze("Sepal.Length", mean)
tbl <- build_table(lyt, iris)
export_tfl(tbl, preview = TRUE)
Top-left label
lyt <- basic_table(title = "Top-Left Label") |>
split_cols_by("Species") |>
append_topleft("Measurement") |>
analyze(c("Sepal.Length", "Petal.Length"), mean)
tbl <- build_table(lyt, iris)
export_tfl(tbl, preview = TRUE)