Creating Tables with tfl_table
v02-tfl_table_intro.Rmdwritetfl can render data frames as paginated,
publication-quality tables inside multi-page PDFs. The two key functions
are tfl_table(), which builds a table configuration object
from a data frame, and export_tfl(), which renders it to a
PDF file.
If you are working with gt tables, see
vignette("v05-gt_tables") for direct gt integration via
export_tfl().
library(writetfl)
library(dplyr) # for group_by() examples
#>
#> Attaching package: 'dplyr'
#> The following objects are masked from 'package:stats':
#>
#> filter, lag
#> The following objects are masked from 'package:base':
#>
#> intersect, setdiff, setequal, unionBasic usage
Pass any data frame to tfl_table(), then hand the result
to export_tfl(). Page annotations, page dimensions, and the
output file all belong to export_tfl() — not to
tfl_table().
tbl <- tfl_table(head(mtcars, 20))
export_tfl(
tbl,
preview = TRUE,
header_left = "Table 1. Motor Trend Car Road Tests (first 20 rows)",
footer_left = "Source: Motor Trend (1974)",
header_rule = TRUE
)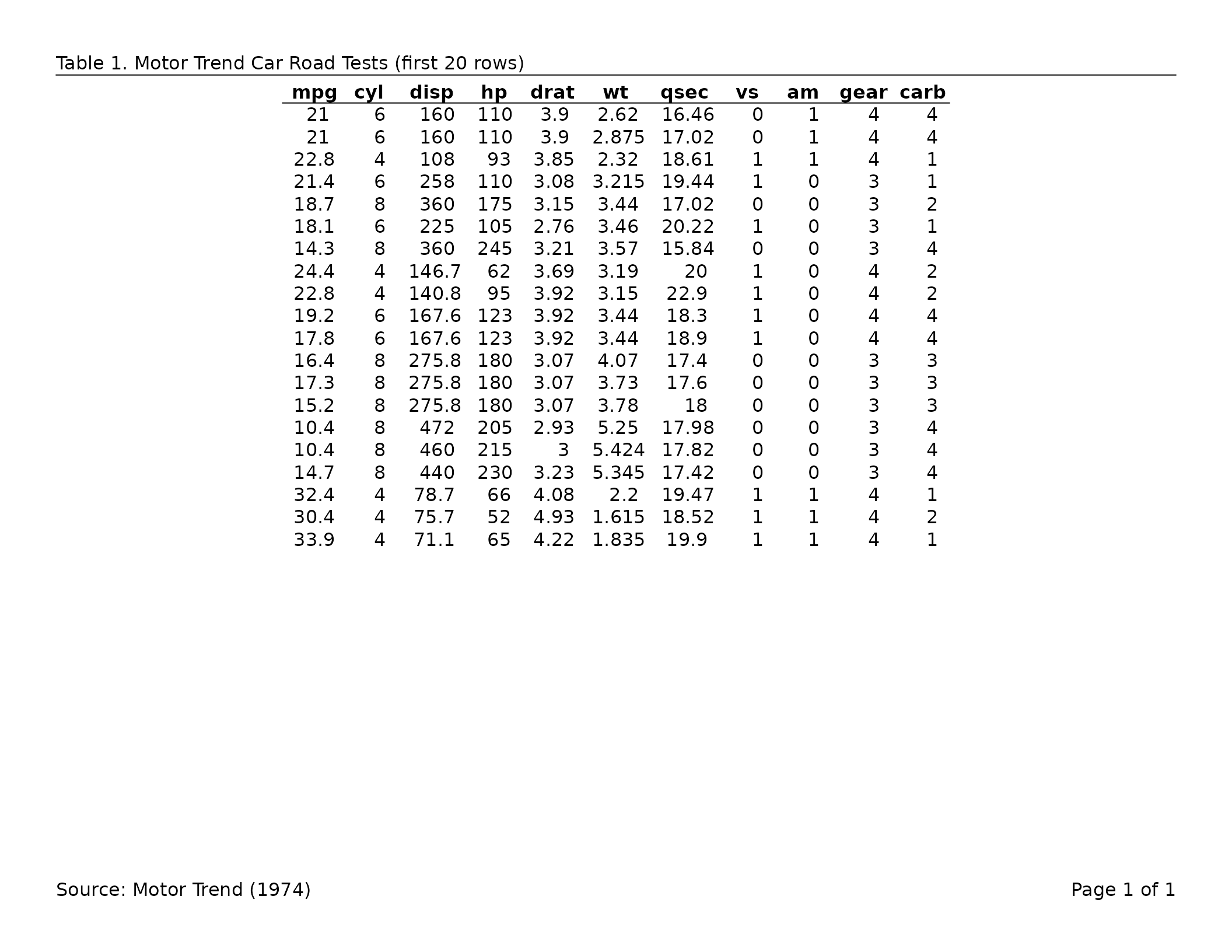
tfl_table() returns a table configuration object. It
does not draw anything or open any device. All drawing happens inside
export_tfl().
Columns
Labels
By default, column names are used as column headers. Supply
col_labels to override them — either as a named character
vector (match by column name) or as a positional vector the same length
as cols.
# Subset columns and rename them for the report
tbl <- tfl_table(
head(mtcars, 20)[, c("mpg", "cyl", "hp", "wt")],
col_labels = c(
mpg = "Miles/Gallon",
cyl = "Cylinders",
hp = "Horsepower",
wt = "Weight\n(1000 lb)" # \n produces a two-line header
)
)
export_tfl(
tbl,
preview = TRUE,
header_left = "Table 1. Selected Performance Metrics"
)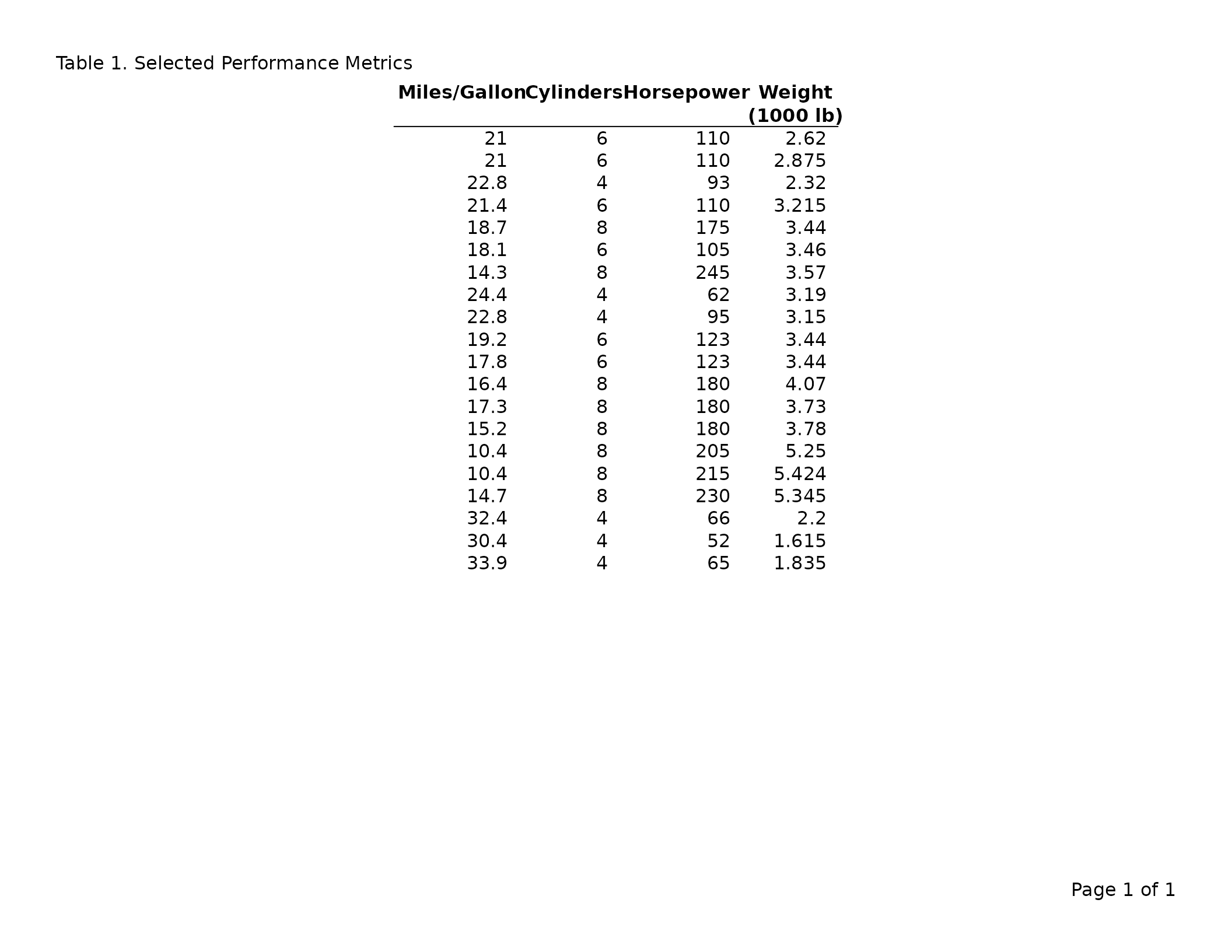
Embedding \n in a label creates a multi-line column
header. The header row height is sized automatically to fit the tallest
label.
Widths
Three width modes are available and can be mixed freely within the same table.
| Mode | How to specify | Effect |
|---|---|---|
| Fixed | unit(1.5, "inches") |
Always exactly that width |
| Relative | Plain numeric, e.g. 2
|
Width proportional to remaining space after fixed columns are placed |
| Auto |
NULL (the default) |
Sized to the widest content, up to a per-column maximum |
tbl <- tfl_table(
head(mtcars, 20)[, c("mpg", "cyl", "hp", "wt", "gear")],
col_widths = list(
mpg = unit(1.2, "inches"), # fixed
cyl = 1, # relative (equal share of remaining space)
hp = 1, # relative (equal share)
wt = unit(1.4, "inches"), # fixed
gear = NULL # auto: sized to content
)
)
export_tfl(tbl, file = "column_widths.pdf")When relative widths are used, they are scaled proportionally among
themselves after all fixed and auto columns have claimed their space.
min_col_width (default unit(0.5, "inches")) is
a floor applied to auto-sized columns.
Alignment
Numeric columns default to right-aligned; character columns default
to left-aligned. Override per column with col_align.
ae_summary <- data.frame(
system_organ_class = c("Gastrointestinal", "Nervous system", "Skin"),
n_subjects = c(12L, 7L, 4L),
pct = c(24.0, 14.0, 8.0),
stringsAsFactors = FALSE
)
tbl <- tfl_table(
ae_summary,
col_labels = c(
system_organ_class = "System Organ Class",
n_subjects = "n",
pct = "(%)"
),
col_align = c(
system_organ_class = "left",
n_subjects = "right",
pct = "right"
)
)
export_tfl(
tbl,
preview = TRUE,
header_left = "Table 2. Adverse Events by System Organ Class",
footnote = "Percentages are based on the safety population (N = 50)."
)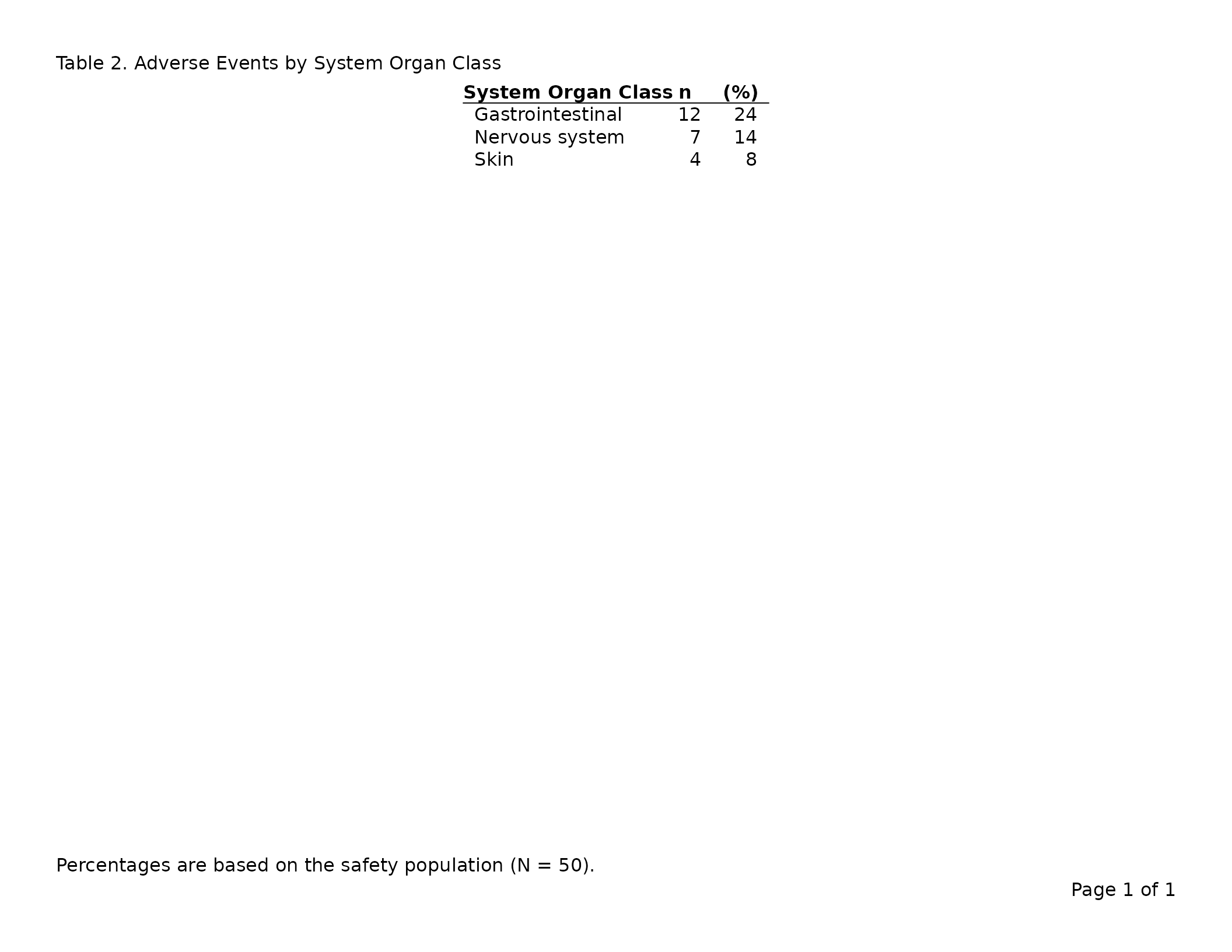
Valid alignment values are "left", "right",
and "center".
Word-wrapping
writetfl ships with text wrapping turned on by default.
wrap_cols controls which columns are eligible:
wrap_cols value |
Meaning |
|---|---|
"auto" (default) |
Eligible if the column’s data or header contains a break character (whitespace by default). Numeric and single-token columns are skipped automatically. |
TRUE |
All non-group columns are eligible regardless of content. |
FALSE |
Disable text wrapping entirely. |
| Character vector of column names | Only the named columns are eligible. |
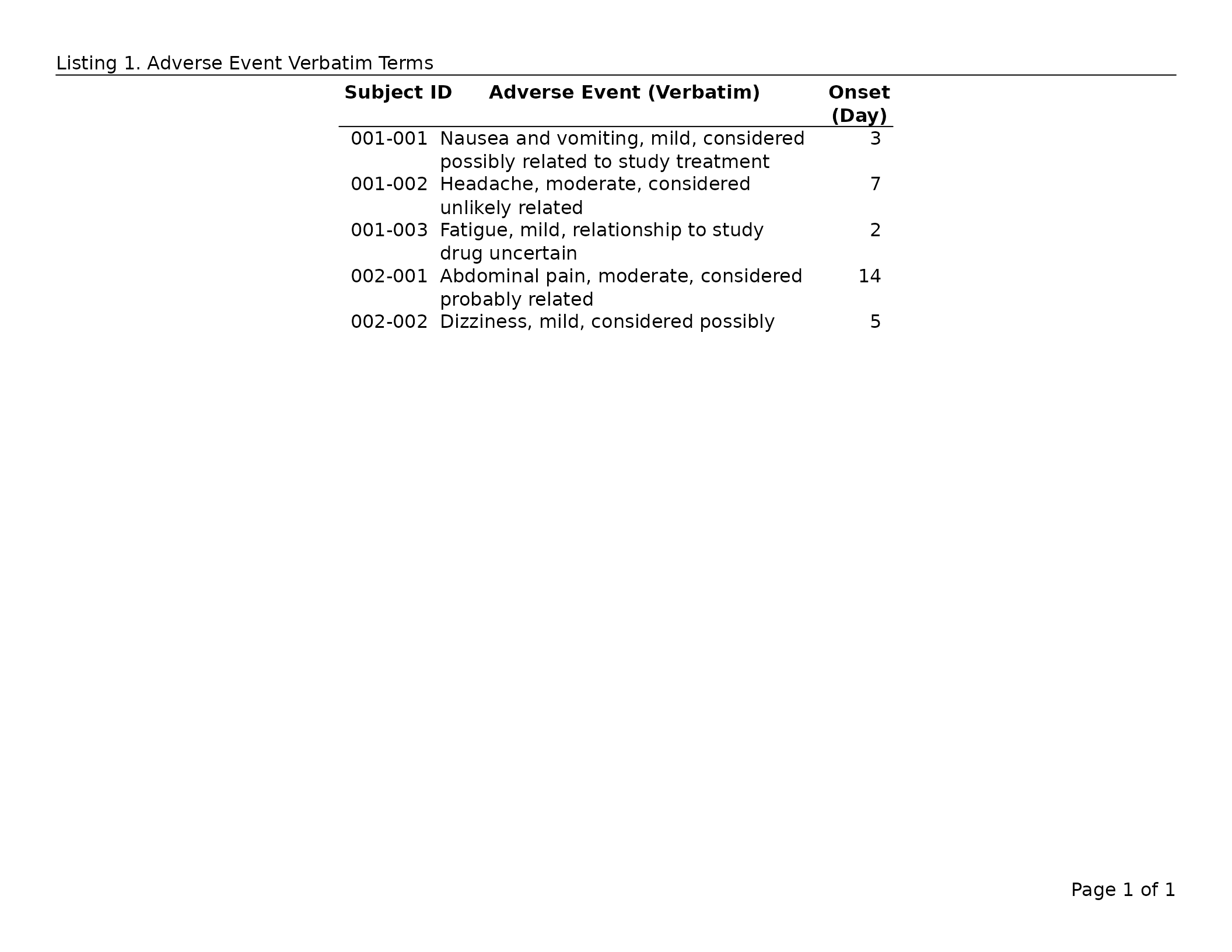
The default (“auto”) is the recommended setting for clinical reports — free-text columns (narrative descriptions, verbatim terms) wrap to fit their column without you having to specify; numeric columns are left at their natural width.
ae_verbatim <- data.frame(
subject_id = c("001-001", "001-002", "001-003", "002-001", "002-002"),
ae_term = c(
"Nausea and vomiting, mild, considered possibly related to study treatment",
"Headache, moderate, considered unlikely related",
"Fatigue, mild, relationship to study drug uncertain",
"Abdominal pain, moderate, considered probably related",
"Dizziness, mild, considered possibly related"
),
onset_day = c(3L, 7L, 2L, 14L, 5L),
stringsAsFactors = FALSE
)
# wrap_cols = "auto" is the default; specifying it is shown for clarity.
tbl <- tfl_table(
ae_verbatim,
col_labels = c(
subject_id = "Subject ID",
ae_term = "Adverse Event (Verbatim)",
onset_day = "Onset\n(Day)"
),
col_widths = list(
subject_id = unit(0.8, "inches"),
ae_term = unit(3.5, "inches"),
onset_day = NULL
)
)
export_tfl(
tbl,
preview = TRUE,
header_left = "Listing 1. Adverse Event Verbatim Terms",
header_rule = TRUE
)
Per-column override
Set tfl_colspec(wrap = TRUE) or
tfl_colspec(wrap = FALSE) to override the table-level
setting for one column. wrap = NA (the default) means
“inherit from wrap_cols”.
Custom break characters — wrap_breaks()
By default the wrap algorithm breaks on whitespace and consumes the
whitespace at the break. wrap_breaks() lets you opt in to
break-after characters that stay on the upper line:
# Break after hyphens for hyphenated terms like "placebo-controlled"
tbl <- tfl_table(
hyphen_df,
wrap_breaks = wrap_breaks(drop = " ", keep_before = "-")
)
# Break after "/" or "-" for path-like content
tbl <- tfl_table(
path_df,
wrap_breaks = wrap_breaks(drop = " ", keep_before = c("-", "/"))
)Optimising for total table height — wrap_balance
The default wrap_balance = "width" keeps the widths of
wrap-eligible columns balanced (water-from-top), which is fast and
produces visually tidy columns. When two wrap-eligible columns have very
different content density, the dense one wraps to many more lines than
the sparse one even at the same width — and the resulting tall rows can
spill across pages. Set wrap_balance = "height" to opt in
to a bounded extra pass that redistributes width between wrap-eligible
columns to lower the total table height (more rows per page). The pass
is time-budgeted and falls back silently to the default if it can’t find
an improvement.
Row-overflow guard
A row whose wrapped height exceeds one page is almost always a sign
of input that needs to change (e.g. a 5,000-character cell forced into a
0.5-inch column). The package raises a clear error in this case;
overflow_action = "warn" on export_tfl()
downgrades it to a warning and produces output for diagnosis. See
vignette("v04-troubleshooting") for the full overflow
story.
Per-column specification with tfl_colspec()
For complex tables it can be cleaner to specify each column
separately using tfl_colspec() and collect the results into
a list. This avoids long parallel vectors for labels, widths, and
alignments.
pk_summary <- data.frame(
param = rep(c("Cmax", "AUC0-inf", "t1/2"), each = 3),
treatment = rep(c("Placebo", "Active 10 mg", "Active 20 mg"), 3),
geo_mean = c(0.00, 145.2, 210.8, 0.00, 4820, 7340, 0.00, 8.4, 9.1),
cv_pct = c(NA, 28.4, 31.2, NA, 22.7, 25.8, NA, 15.3, 17.9),
stringsAsFactors = FALSE
)
tbl <- pk_summary |>
group_by(param) |>
tfl_table(
cols = list(
tfl_colspec("param", label = "Parameter", width = unit(1.2, "inches"), align = "left"),
tfl_colspec("treatment", label = "Treatment", width = unit(1.5, "inches"), align = "left"),
tfl_colspec("geo_mean", label = "Geometric\nMean", width = unit(1.2, "inches"), align = "right"),
tfl_colspec("cv_pct", label = "CV%", width = unit(0.8, "inches"), align = "right")
),
na_string = "--"
)
export_tfl(
tbl,
preview = TRUE,
header_left = "Table 3. PK Parameters — Geometric Mean (CV%)",
footnote = c(
"CV% = coefficient of variation.",
"-- = not applicable (placebo)."
)
)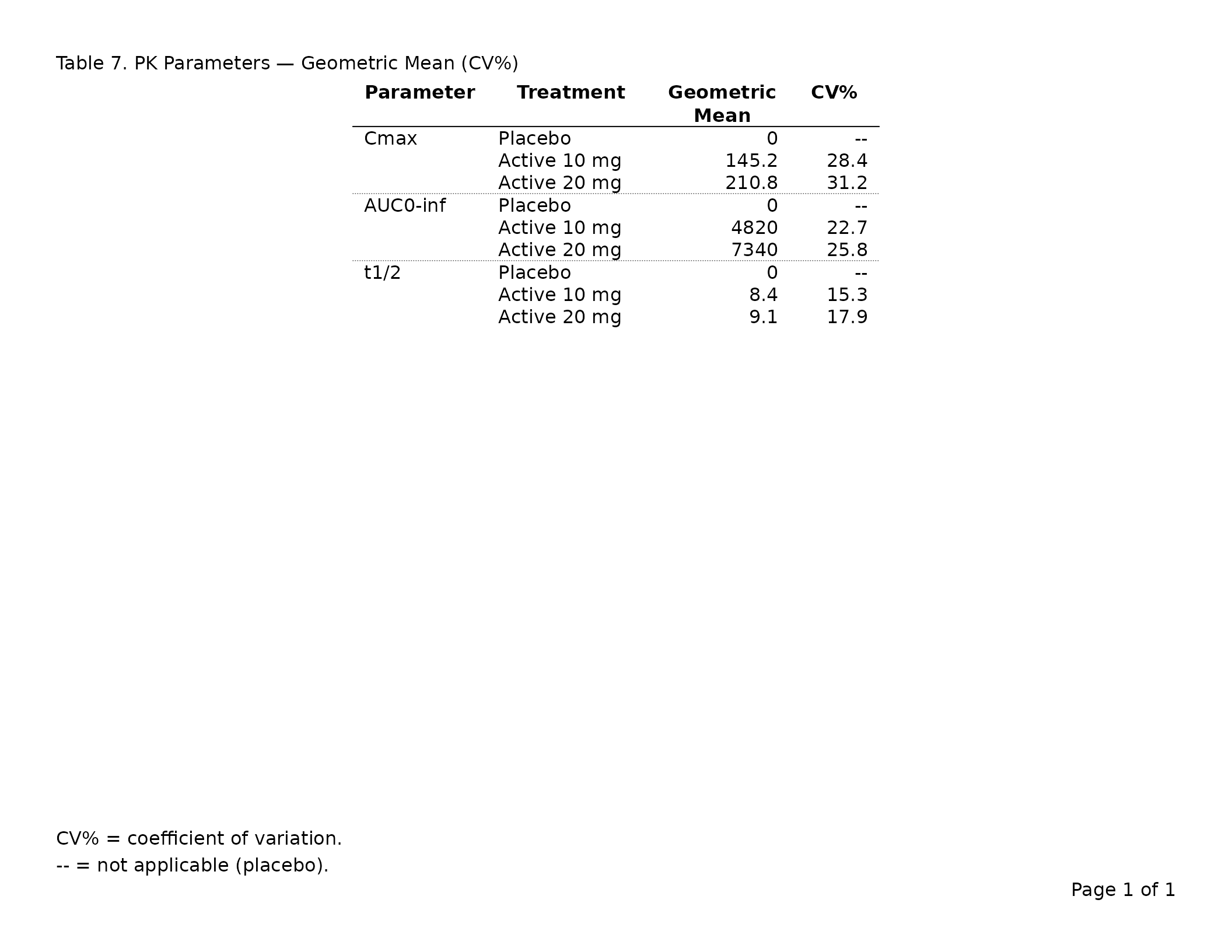
tfl_colspec() accepts col,
label, width, align,
wrap, and gp. It provides no functionality
beyond what the parallel-vector approach offers; the choice is
stylistic.
Rows
Row grouping
Mark grouping columns with dplyr::group_by() before
passing the data to tfl_table(). Group columns must appear
first in the column list; they act as row headers and their values are
suppressed on repeated consecutive rows, giving the indented-group
appearance common in clinical tables.
# Demographic summary with visit and treatment group as row headers
pk_data <- data.frame(
visit = rep(c("Week 4", "Week 8", "Week 12"), each = 4),
treatment = rep(c("Placebo", "Active 10 mg", "Active 20 mg", "Active 40 mg"), 3),
n = c(48L, 50L, 49L, 51L, 45L, 47L, 48L, 50L, 41L, 43L, 44L, 46L),
mean_auc = c(120.4, 145.2, 178.9, 201.3,
118.7, 148.6, 185.2, 219.4,
115.1, 152.3, 191.7, 228.6),
sd_auc = c(18.2, 22.4, 27.6, 31.1,
17.9, 23.1, 28.4, 32.7,
17.1, 24.0, 29.2, 34.3),
stringsAsFactors = FALSE
)
tbl <- pk_data |>
group_by(visit) |>
tfl_table(
col_labels = c(
visit = "Visit",
treatment = "Treatment",
n = "n",
mean_auc = "Mean AUC\n(ng·h/mL)",
sd_auc = "SD"
),
col_widths = list(
visit = unit(1.0, "inches"),
treatment = unit(1.5, "inches"),
n = NULL,
mean_auc = unit(1.4, "inches"),
sd_auc = unit(0.8, "inches")
)
)
export_tfl(
tbl,
preview = TRUE,
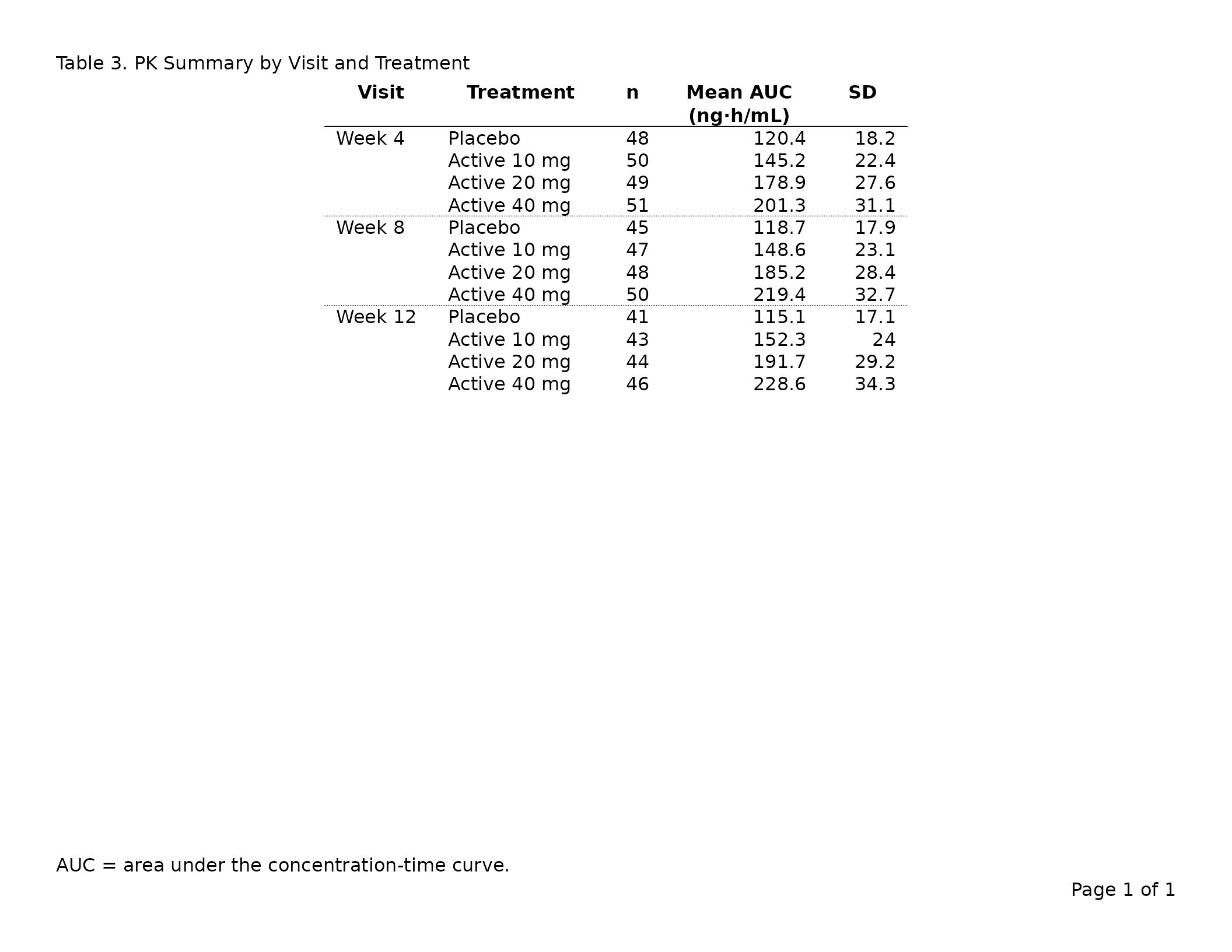
header_left = "Table 4. PK Summary by Visit and Treatment",
footnote = "AUC = area under the concentration-time curve."
)
Group values within a block are kept together on the same page wherever possible (see Pagination below). Multiple group columns are honored — the leftmost is the outermost grouping, with each subsequent column nested inside its parent.
Suppression and multi-line group labels
suppress_repeated_groups (default TRUE)
blanks group-column cells whose value matches the previous rendered row.
The first row on each page always shows the group value, so a reader can
identify the group of any visible block of rows.
When the group label itself spans multiple lines (because of an
embedded \n or word-wrapping), the label flows downward
through the blanked cells like an HTML
<td rowspan="N"> instead of inflating only the
labelled row. The example below has a two-line group label
(“TreatmentA”) that flows across all three rows of the group; no row
needs to grow taller than its data content.
flow_demo <- data.frame(
group = rep("Treatment\nArm A", 3L),
visit = c("Day 1", "Day 8", "Day 15"),
value = c("12.3", "15.7", "18.1"),
stringsAsFactors = FALSE
) |> group_by(group)
export_tfl(
tfl_table(flow_demo),
preview = TRUE,
header_left = "Multi-line group label flows across rows"
)
Two consequences flow from this layout choice:
- Row heights are computed per page. The same data row may render at different heights on different pages when a group is split: the first row on each page re-shows the label and may need to grow alone if the rest of the group landed on a different page.
-
Rules adapt to the flow. Row rules (see
vignette("v03-tfl_table_styling")) are suppressed inside a multi-row group span so the flowing label isn’t visually sliced. Group rules draw at every transition and start at the column whose value actually changed at that boundary, so unchanged outer columns through which the label is flowing aren’t visually divided.
If you would rather render every group cell explicitly on every row
(no flow, no blanking), set
suppress_repeated_groups = FALSE. Each row’s height then
becomes the per-row max over every cell, including the multi-line group
label.
Missing values
na_string controls how NA values are
displayed in the table. The default is "" (an empty cell).
Supply any character string to substitute a visible token.
labs_data <- data.frame(
subject_id = c("001", "001", "002", "002", "003"),
visit = c("Baseline", "Week 4", "Baseline", "Week 4", "Baseline"),
ALT = c(28, 31, NA, 45, 22),
AST = c(19, NA, 24, 38, 17),
stringsAsFactors = FALSE
)
tbl <- labs_data |>
group_by(subject_id) |>
tfl_table(
col_labels = c(
subject_id = "Subject",
visit = "Visit",
ALT = "ALT\n(U/L)",
AST = "AST\n(U/L)"
),
na_string = "NC" # NC = not collected
)
export_tfl(
tbl,
preview = TRUE,
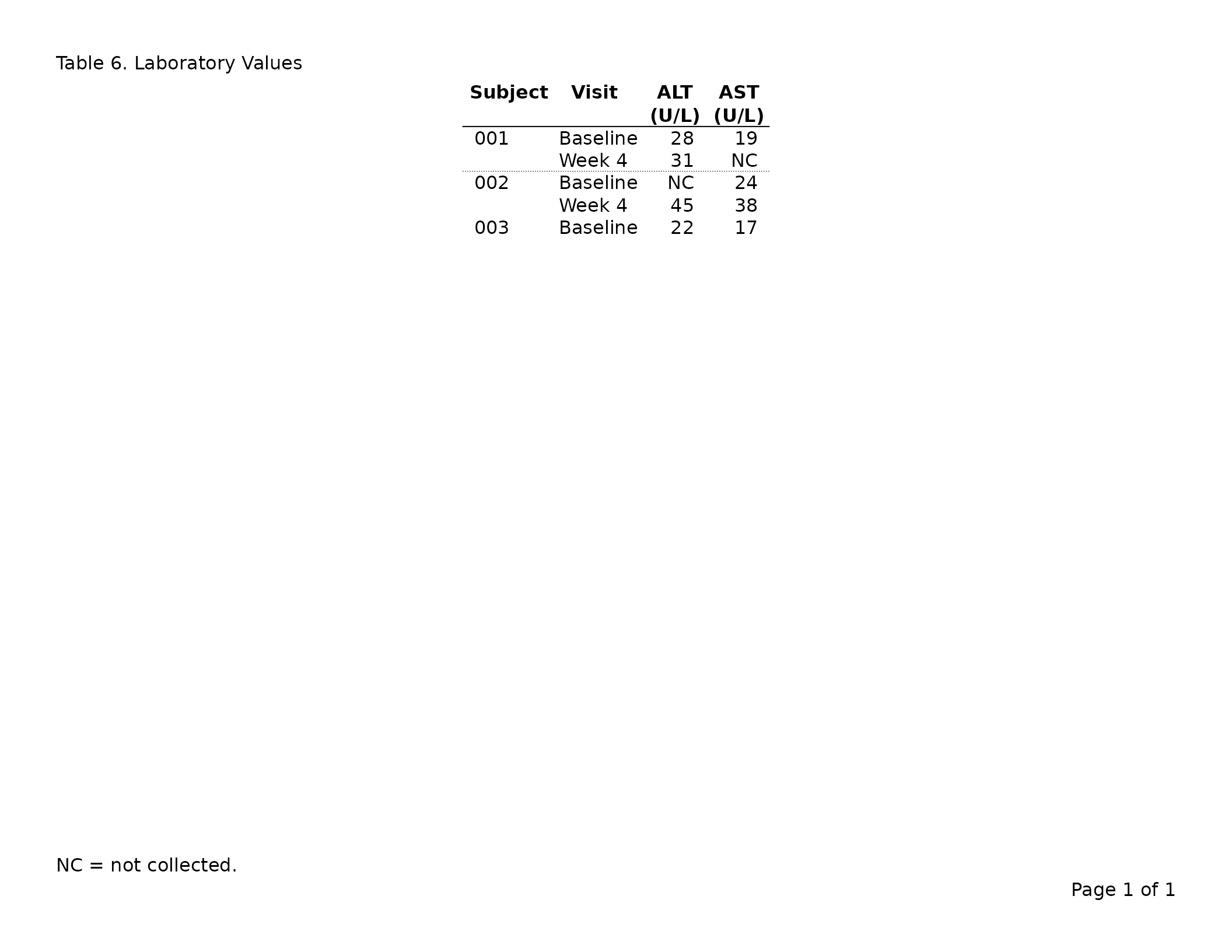
header_left = "Table 5. Laboratory Values",
footnote = "NC = not collected."
)
Pagination
Row pagination
When a table has more rows than fit on one page,
tfl_table paginates automatically. Groups are kept
together: if the rows belonging to one group value do not fit on the
current page, the whole group moves to the next page wherever
possible.
When a group genuinely cannot fit on a single page, it is split, and continuation markers are appended to the last column header on intermediate pages and to the first data row of a continuation page so the reader can follow the table across page breaks.
tbl <- iris |>
relocate(Species) |>
group_by(Species) |>
tfl_table(
col_labels = c(
Species = "Species",
Sepal.Length = "Sepal\nLength",
Sepal.Width = "Sepal\nWidth",
Petal.Length = "Petal\nLength",
Petal.Width = "Petal\nWidth"
),
row_cont_msg = c("(continued from previous page)", "(continued on next page)"),
col_cont_msg = "(continued)"
)
export_tfl(
tbl,
preview = c(1, 2),
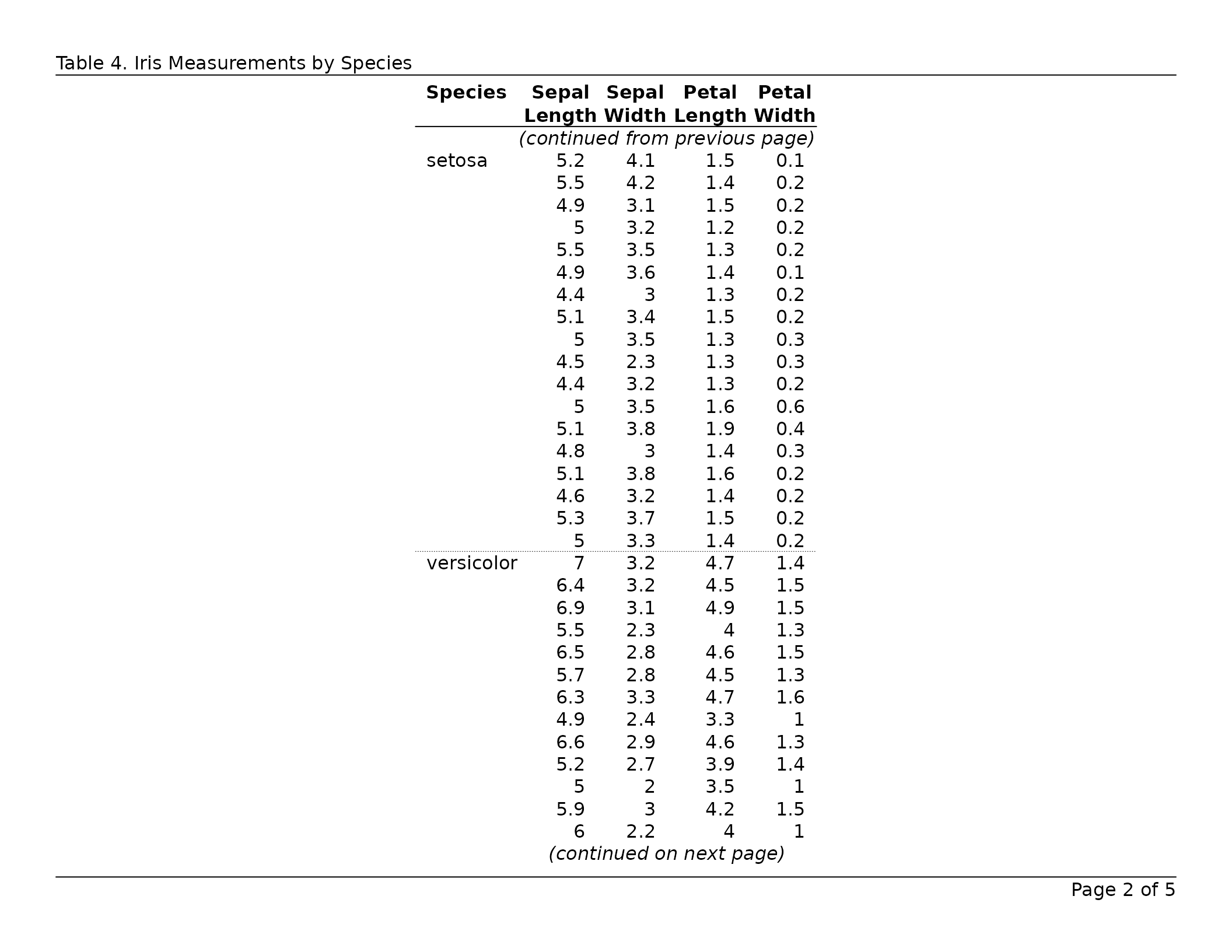
header_left = "Table 6. Iris Measurements by Species",
header_rule = TRUE,
footer_rule = TRUE
)
#> Warning: Row 32 belongs to a group that spans more than one page. A
#> '(continued)' marker will be added at the boundary.
#> Warning: Row 62 belongs to a group that spans more than one page. A
#> '(continued)' marker will be added at the boundary.
#> Warning: Row 92 belongs to a group that spans more than one page. A
#> '(continued)' marker will be added at the boundary.
#> Warning: Row 122 belongs to a group that spans more than one page. A
#> '(continued)' marker will be added at the boundary.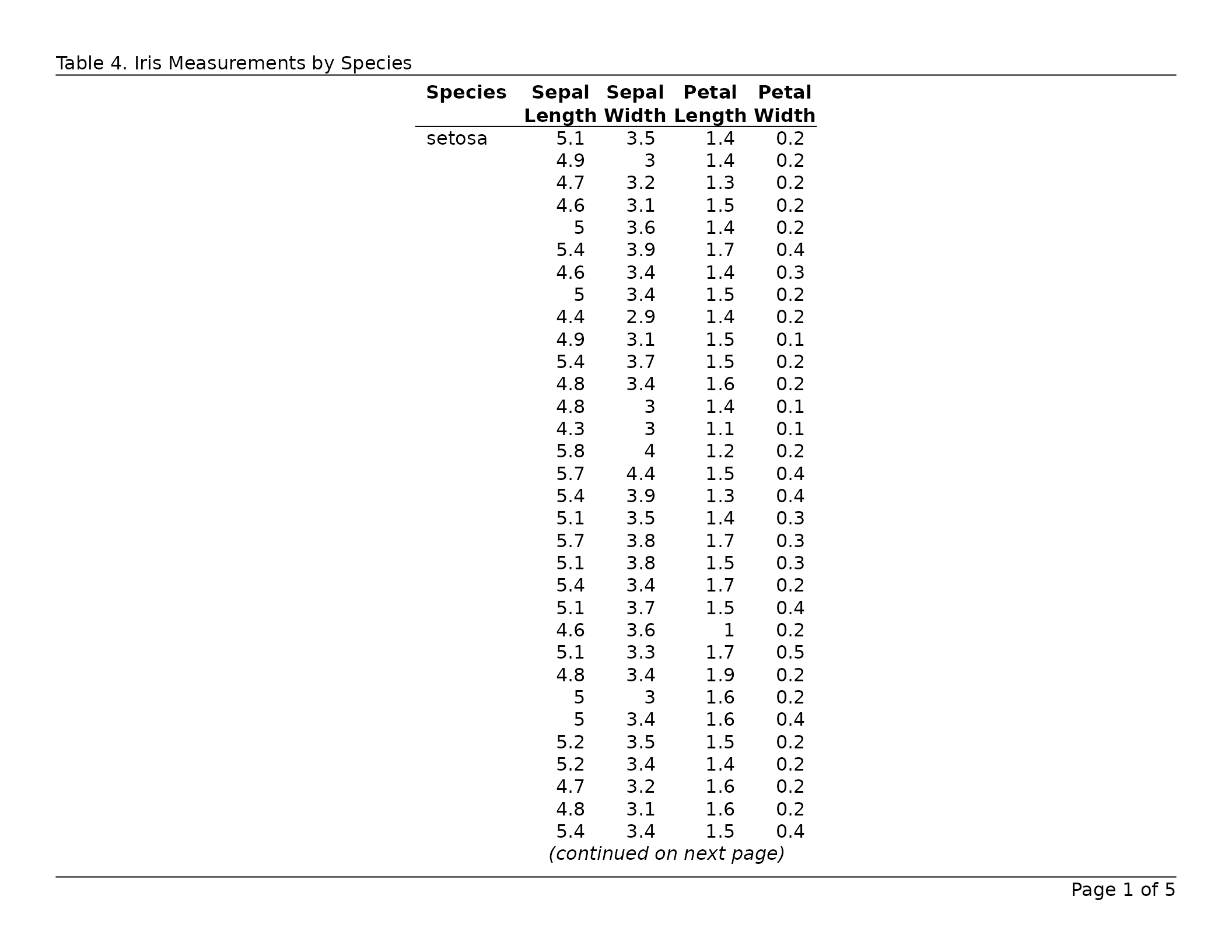
Column pagination
If the total column width exceeds the printable area,
tfl_table splits the columns across additional pages.
Row-header columns (i.e., the grouped columns) are repeated at the left
of every column page so the reader always knows which group they are
in.
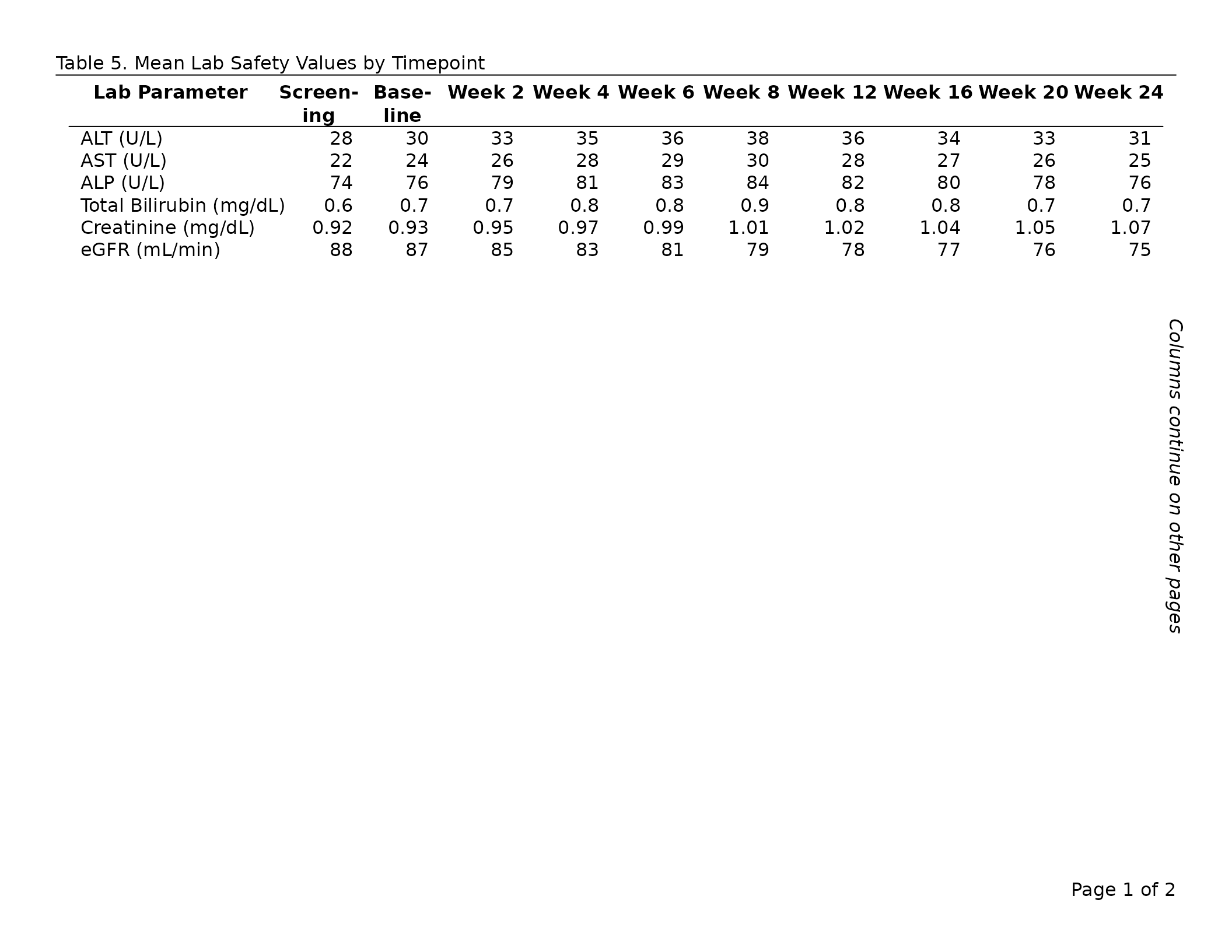
# Lab safety panel: 6 parameters × 13 timepoints — too wide for one page.
# Group by parameter so the parameter column repeats as a row-header on each
# column-split page.
lab_wide <- data.frame(
parameter = c("ALT (U/L)", "AST (U/L)", "ALP (U/L)",
"Total Bilirubin (mg/dL)", "Creatinine (mg/dL)", "eGFR (mL/min)"),
scr = c(28, 22, 74, 0.6, 0.92, 88),
bl = c(30, 24, 76, 0.7, 0.93, 87),
wk2 = c(33, 26, 79, 0.7, 0.95, 85),
wk4 = c(35, 28, 81, 0.8, 0.97, 83),
wk6 = c(36, 29, 83, 0.8, 0.99, 81),
wk8 = c(38, 30, 84, 0.9, 1.01, 79),
wk12 = c(36, 28, 82, 0.8, 1.02, 78),
wk16 = c(34, 27, 80, 0.8, 1.04, 77),
wk20 = c(33, 26, 78, 0.7, 1.05, 76),
wk24 = c(31, 25, 76, 0.7, 1.07, 75),
wk28 = c(30, 24, 75, 0.6, 1.08, 74),
wk32 = c(29, 23, 74, 0.6, 1.09, 73),
eot = c(28, 22, 73, 0.6, 1.10, 72),
stringsAsFactors = FALSE
)
tbl <- lab_wide |>
group_by(parameter) |>
tfl_table(
col_labels = c(
parameter = "Lab Parameter",
scr = "Screen-\ning", bl = "Base-\nline",
wk2 = "Week 2", wk4 = "Week 4",
wk6 = "Week 6", wk8 = "Week 8",
wk12 = "Week 12", wk16 = "Week 16",
wk20 = "Week 20", wk24 = "Week 24",
wk28 = "Week 28", wk32 = "Week 32",
eot = "End of\nTreatment"
)
)
export_tfl(
tbl,
preview = c(1, 2),
header_left = "Table 7. Mean Lab Safety Values by Timepoint",
header_rule = TRUE
)
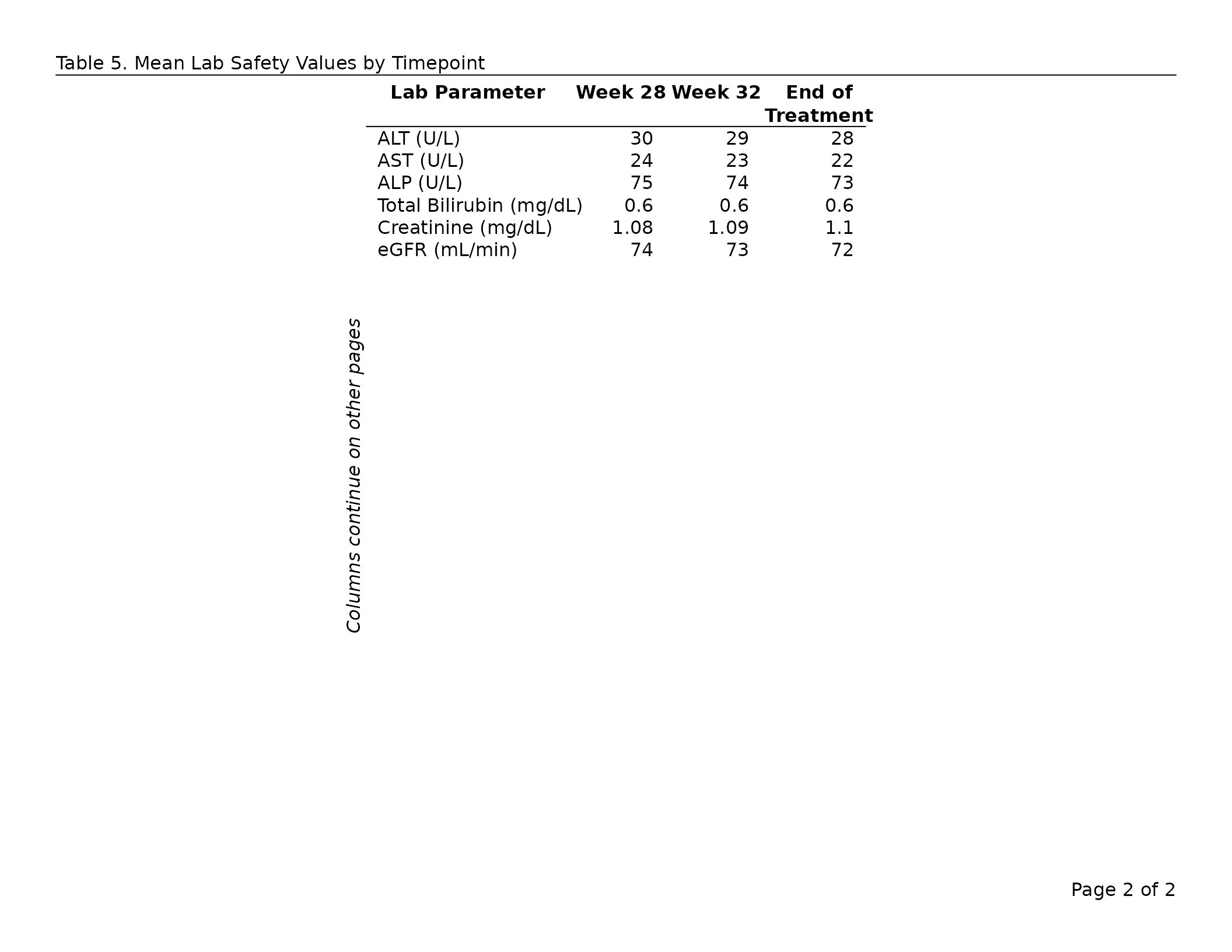
By default allow_col_split = TRUE. Set it to
FALSE if you want an error rather than an automatic split —
useful during development to confirm that your column widths fit within
the target page dimensions.
# This will error if the columns are too wide for the page
tbl_no_split <- tfl_table(
mtcars,
allow_col_split = FALSE
)
export_tfl(tbl_no_split, preview = TRUE)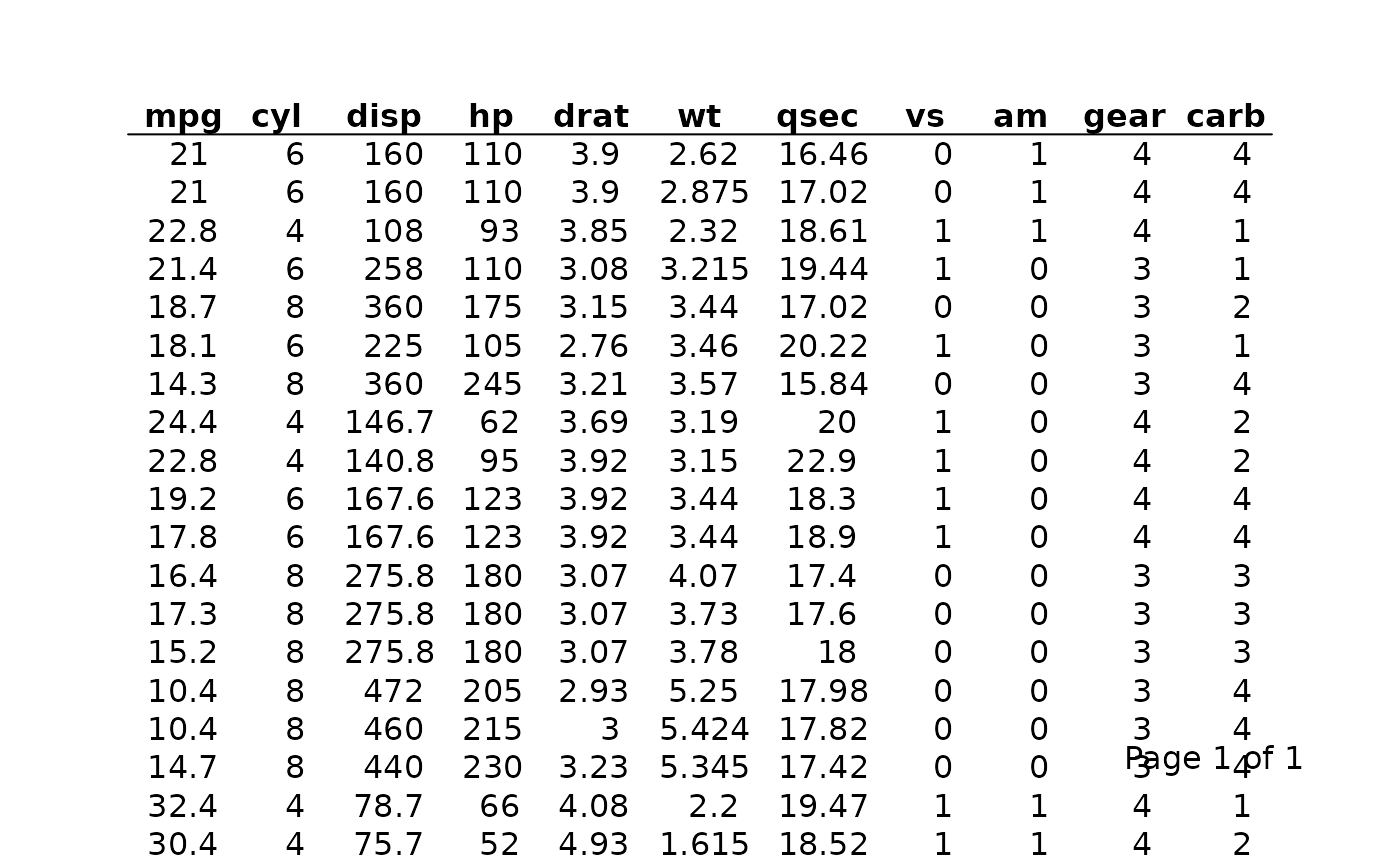
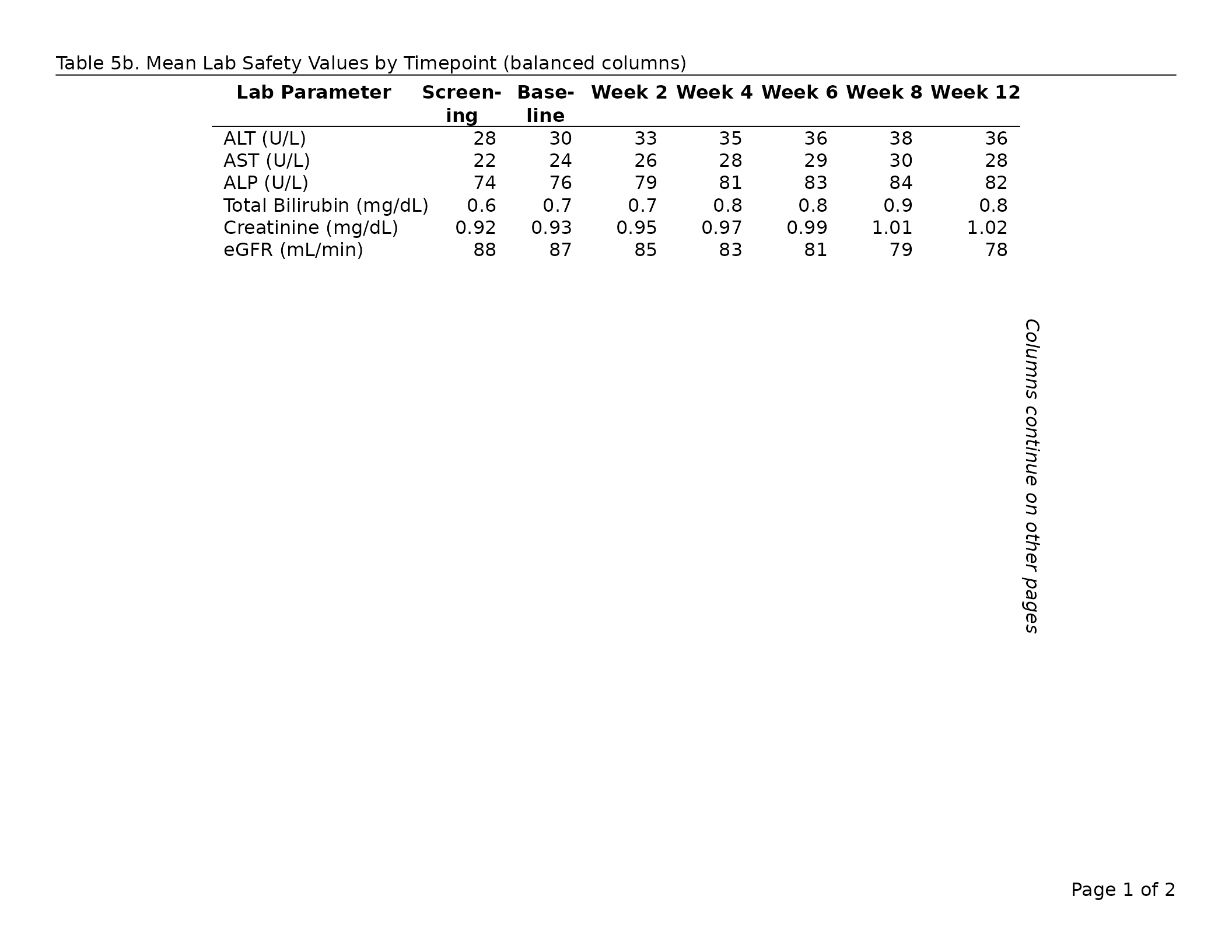
Balancing columns across pages
By default the greedy algorithm packs as many columns as possible
onto each page, which can leave the last page looking sparse — for
example, 8 columns on page 1 and only 2 on page 2. Set
balance_col_pages = TRUE to redistribute the data columns
so that each page carries approximately the same number.
The greedy pass still runs first to determine the minimum number of pages required. The data columns are then divided as evenly as possible across those pages (pages that cannot be divided exactly get one extra column on the earlier pages). Each candidate balanced group is verified to fit within the available width; if any group would overflow, the greedy layout is used as a fallback.
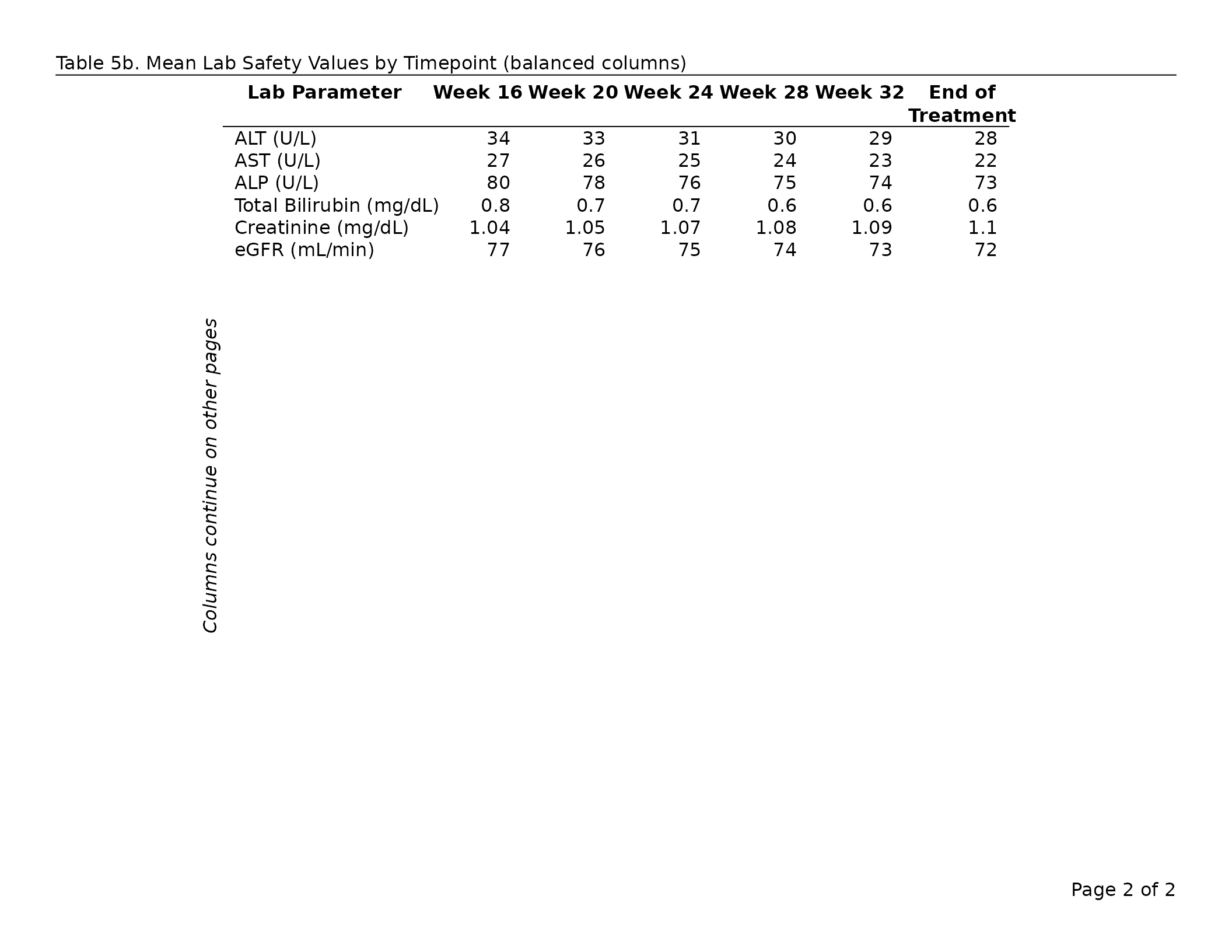
tbl_balanced <- lab_wide |>
group_by(parameter) |>
tfl_table(
col_labels = c(
parameter = "Lab Parameter",
scr = "Screen-\ning", bl = "Base-\nline",
wk2 = "Week 2", wk4 = "Week 4",
wk6 = "Week 6", wk8 = "Week 8",
wk12 = "Week 12", wk16 = "Week 16",
wk20 = "Week 20", wk24 = "Week 24",
wk28 = "Week 28", wk32 = "Week 32",
eot = "End of\nTreatment"
),
balance_col_pages = TRUE
)
export_tfl(
tbl_balanced,
preview = c(1, 2),
header_left = "Table 7b. Mean Lab Safety Values (balanced columns)",
header_rule = TRUE
)
Page layout and annotations
Page dimensions, margins, header/footer text, separator rules, and
page numbering are all arguments to export_tfl(), not
tfl_table(). This keeps the table structure independent
from the output format, allowing the same tfl_table object
to be used with different page layouts.
tbl <- tfl_table(
head(iris, 30),
col_labels = c(
Species = "Species",
Sepal.Length = "Sepal\nLength",
Sepal.Width = "Sepal\nWidth",
Petal.Length = "Petal\nLength",
Petal.Width = "Petal\nWidth"
)
)
export_tfl(
tbl,
preview = TRUE,
pg_width = 8.5,
pg_height = 11,
margins = unit(c(t = 1, r = 0.75, b = 1, l = 0.75), "inches"),
header_left = "Protocol XY-001\nDraft — Not for Distribution",
header_center = "CONFIDENTIAL",
header_right = format(Sys.Date(), "%d %b %Y"),
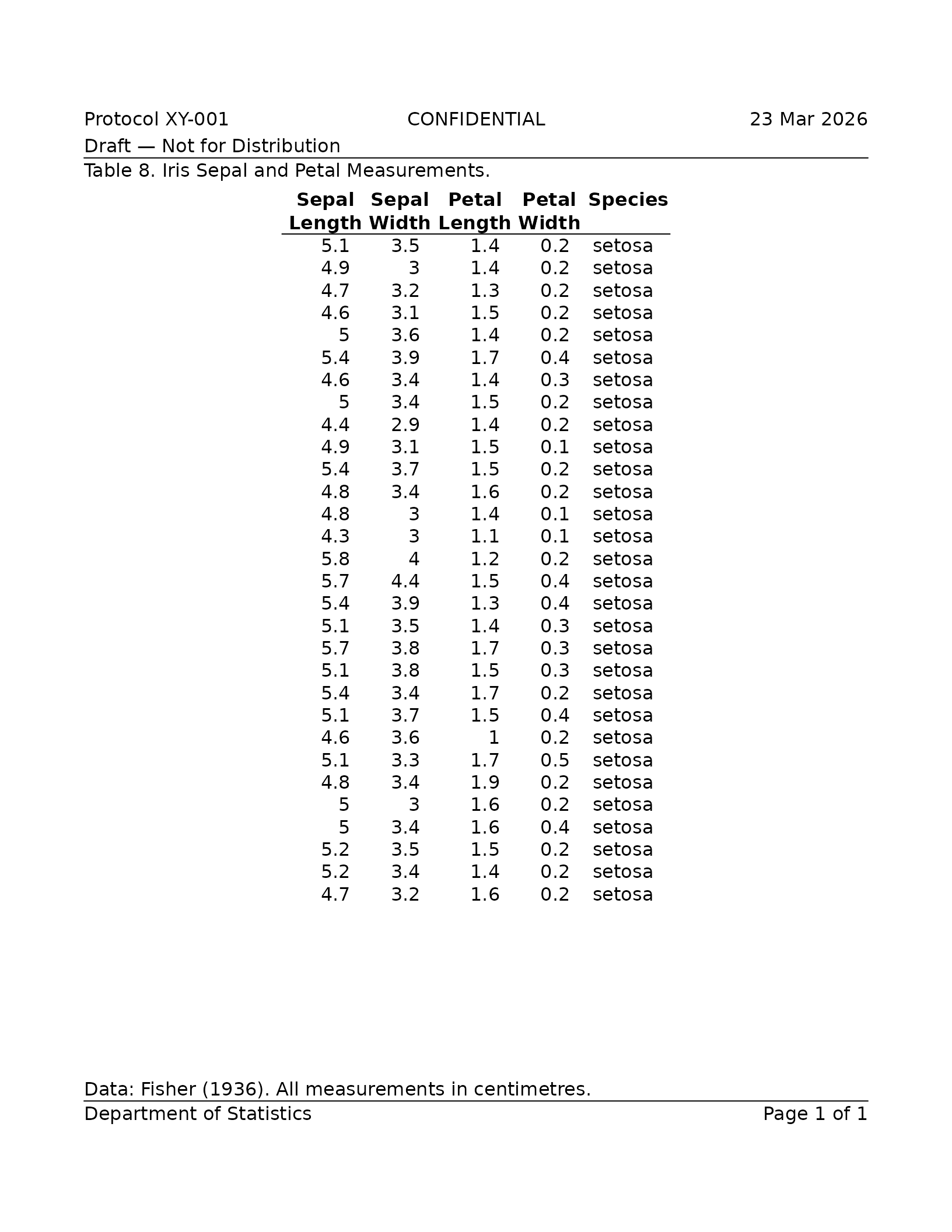
caption = "Table 8. Iris Sepal and Petal Measurements.",
footnote = "Data: Fisher (1936). All measurements in centimetres.",
footer_left = "Department of Statistics",
header_rule = TRUE,
footer_rule = TRUE
)
See vignette("v01-figure_output") for a full reference
of all export_tfl() layout arguments, including typography
(gp), padding, rules, and overlap detection.
Typography (preview)
The gp argument to tfl_table() controls
cell typography. Pass a single gpar() for a uniform style,
or a named list for per-section control.
tbl <- tfl_table(
head(mtcars, 15)[, c("mpg", "cyl", "hp", "wt")],
col_labels = c(mpg = "MPG", cyl = "Cylinders", hp = "Horsepower", wt = "Weight"),
gp = list(
table = gpar(fontsize = 9),
header_row = gpar(fontface = "bold")
)
)For a full reference of all gp keys and their effects
(typography, rules, fills, sub-tables, and the complete list of styling
arguments), see vignette("v03-tfl_table_styling").
Summary of tfl_table() arguments
| Argument | Default | Purpose |
|---|---|---|
x |
— | Data frame or grouped tibble |
| Columns | ||
cols |
NULL (all columns) |
NULL or a list of tfl_colspec() objects.
To display a column subset, pre-select columns in x before
passing to tfl_table(). |
col_widths |
NULL (auto) |
Named list of unit(), plain numeric, or
NULL per column |
col_labels |
column names | Named character vector of header labels; supports
\n
|
col_align |
type-based | Named vector: "left", "right", or
"center"
|
wrap_cols |
"auto" |
"auto" (auto-detect), TRUE (all data
cols), FALSE (off), or character vector of column
names |
wrap_breaks |
wrap_breaks() |
Break-character spec — defaults to whitespace; opt into
keep_before chars like - or
/
|
wrap_balance |
"width" |
"width" (fast water-fill) or "height"
(opt-in pass that lowers total table height) |
col_split_strategy |
"balanced" |
"balanced" (page-split using min widths, then
water-fill per page — more horizontal room per column on multi-page
tables) or "wrap_first" (legacy: whole-table water-fill,
then split using post-wrap widths) |
row_overflow_max_retries |
5L |
When a row’s wrapped height exceeds the page under
col_split_strategy = "balanced", raise the bottleneck
column’s minimum by 0.25 in and retry up to N times. 0L
disables the retry loop. |
min_col_width |
unit(0.5, "inches") |
Floor applied to auto-sized columns |
allow_col_split |
TRUE |
If FALSE, error when columns exceed page width |
balance_col_pages |
FALSE |
Redistribute columns evenly across column-split pages |
| Rows and grouping | ||
suppress_repeated_groups |
TRUE |
Blank repeated group values; multi-line labels flow into the blanked cells (HTML-rowspan style) |
na_string |
"" |
Replacement for NA values |
| Pagination | ||
col_cont_msg |
side labels | Rotated text on column-split pages |
row_cont_msg |
c("(continued)", "(continued on next page)") |
[1] shown at top of continuation page; [2]
shown at bottom of page before continuation |
| Rules and separators | ||
show_col_names |
TRUE |
Whether to render the column header row at all |
col_header_rule |
TRUE |
Rule below column headers |
group_rule |
TRUE |
Rule at each new group block (partial width — see
vignette("v03-tfl_table_styling")) |
group_rule_after_last |
FALSE |
Rule after the last group block |
row_rule |
FALSE |
Rule between data rows (suppressed inside multi-row group spans) |
row_header_sep |
FALSE |
Vertical rule after row-header columns |
| Sub-tables | ||
sub_tfl |
NULL |
One self-identifying sub-table per unique combination of values; see
vignette("v03-tfl_table_styling")
|
sub_tfl_sep / sub_tfl_collapse /
sub_tfl_prefix
|
": " / "; " / "\n"
|
Caption-suffix formatting for sub-tables |
| Typography and spacing | ||
gp |
list() |
Typography for headers and body cells (see
vignette("v03-tfl_table_styling")) |
fill_by |
"row" |
"row" or "group" for cell fill
cycling |
cell_padding |
unit(c(0.2, 0.5), "lines") |
Vertical and horizontal padding inside each cell |
line_height |
1.05 |
Inter-line spacing multiplier for word-wrapped cells |
wrap_extra_padding |
unit(0.5, "lines") |
Extra space below multi-line cells so rows are visually
distinguishable; unit(0, "lines") to disable |
max_measure_rows |
Inf |
Number of rows sampled when measuring auto column widths |